GENETIC_ALGORITHM Function
Optimizes a user-defined fitness function using a tailored genetic algorithm.
Synopsis
result = GENETIC_ALGORITHM ( fitness_fcn, initial_population)
Required Arguments
fitness_fcn—The fitness function. Given the data structure for an individual within the population, fitness returns the fitness of that individual. The fitness function must return non-negative values.
initial_population—A structure containing the initial population.
Return Value
result—GENETIC_ALGORITHM optimizes a user-defined fitness function by evolving an initial population using a tailored genetic algorithm that searches for the fittest individual. It returns a copy of the fittest individual in the last generation.
Input Keywords
Double—If present and nonzero, double precision is used.
Gray_encoding—If present and nonzero, specifies alleles are encoded using Gray encoding for integer and real phenotypes. If undefined or zero, specifies alleles are encoded using Base-2 encoding for integer and real phenotypes. Default: Base-2 encoding.
No_elitism—If present and nonzero, this argument disables elitism. Elitism is used to preserve the fittest individual from one generation to the next. Default: Elitism is enabled.
No_decode—If present and nonzero, this argument disables automatic decoding between generations. By default, chromosome information is decoded into the individual’s phenotypes before every call to the user’s fitness function. Decoding is only applied to the last generation, including the fittest individual. Default: Decoding is enabled.
Print_level—A scalar long value indicating the level of intermediate and final results detail to print. Print_level accepts the following values:
level | Enumeration | Description |
|---|---|---|
0 | IMSLS_NONE | Supresses printing of any results. |
1 | IMSLS_FINAL | Prints summary of final results. |
2 | IMSLS_TRACE_GEN | Prints summary of final results plus generation statistics. |
3 | IMSLS_TRACE_ALL | Prints summary of final results, generation statistics and individual crossovers. |
Default: Print_level = IMSLS_NONE.
Max_generations—A scalar long indicating the maximum number of generations. Optimization is halted when the number of generations exceeds Max_generations. Default: Max_generations=100.
Max_fitness—A scalar float indicating the maximum fitness value. The optimization is halted if the maximum fitness is greater than this value. Default: Max_fitness=MACHINE(POS_INF), i.e., optimization is not halted by large fitness values. Optimization only stops when the number of generations exceeds Max_generations.
Linear_scaling—A scalar float specifying an upper limit for the linear fitness scaling constant. Set Linear_scaling = 1 for no scaling. A check is made to ensure that the minimum scaled fitness is non-negative. If it falls below zero, then the scaling constant is automatically reduced to make the minimum scaled fitness equal to zero. For linear scaling the scaling constant is typically between one and two. Default: Linear_scaling = 1, no linear fitness scaling.
Sigma_scaling—If present and nonzero, this keyword enables sigma scaling. Default: No sigma scaling is performed.
Gen_gap—A scalar float indicating the proportion of weakest individuals replaced between generations. If Gen_gap = 1, all of the individuals are replaced. Default: Gen_gap = 1.
Mutate_prob—A scalar float indicating the probability of mutation. Although most applications set this to a value between 0.005 and 0.1, any value between 0 and 1 is allowed. Default: Mutate_prob=0.005.
Crossover_prob—A scalar float indicating the probability of crossover. Crossover_prob can be any value between 0 and 1. Most genetic algorithms use a probability between 0.6 and 0.9. Default: Crossover_prob= 0.6.
N_crossover_pts—A scalar long indicating the number of crossover points. De Jong’s (1975) generalized crossover model R6 is implemented. If N_crossover_pts is odd, then the chromosome is treated as a string with a default crossover at the beginning of the chromosome. If N_crossover_pts is even, then the chromosome is treated as a ring with no beginning or end, and crossovers are selected using the uniform distribution on a circle. Crossing points occur at the odd crossover points. If the Pmx_crossover keyword is used, there are always two crossover points within the nominal portion of the chromosome. For partially matched crossovers, this argument is only used to define the number of crossovers within the binary, integer and real portion of the chromosome. Default: N_crossover_pts = 1.
Pmx_crossover—If present and nonzero, crossover points are randomly selected separately for nominal and non-nominal alleles. By default this keyword applies partially matched crossover to the nominal portion of the chromosome. Crossovers for other phenotypes are still applied using standard crossover and inversion crossover if requested. With partially matched crossover, the number of crossovers for nominal phenotypes is set to 2, and partially matched crossover is applied only to the nominal phenotype. The number of crossovers for non-nominal phenotypes is still controlled by the value of N_crossover_pts. Default: Partially matched crossover is not used.
Invert_crossover—If present and nonzero, inverts the values of the alleles in every other crossover segment. This augments standard or partially matched crossover with inversion. If this is applied with partially matched crossover, inversion is applied within the matched segment of the alleles for the nominal phenotypes and then within every other segment of any non-nominal phenotype. Default: No inversion is performed.
Selection_model—A scalar long value that sets the model used for selecting individuals for reproduction. Selection models are described in the following table:
Value | Selection_model | Description |
|---|---|---|
0 | IMSLS_ROULETTE_WITH | Original fitness-proportional selected described by Holland(1975). Sampling is done with replacement. |
1 | IMSLS_ROULETTE_WITHOUT | The original fitness-proportional selected except that sampling is done without replacement. This is also referred to as De Jong’s (1975) R3 model. |
2 | IMSLS_DETERMINISTIC | Individuals with highest fitness are selected for reproduction using their expected sampling frequency. See Goldberg (1989). |
3 | IMSLS_REMAINDER_WITH | Remainder selection with replacement. |
4 | IMSLS_REMAINDER_WITHOUT | Remainder selection without replacement |
5 | IMSLS_SUS_SELECTION | Stochastic Universal Sampling as described by Baker (1987). |
6 | IMSLS_RANK_SELECTION | Rank selection. The individuals with the highest fitness are selected once for reproduction. |
7 | IMSLS_TOURNAMENT_1 | Tournament selection as described by Wetzel (1983). Only the fittest individual in a pair is selected. |
8 | IMSLS_TOURNAMENT_2 | Tournament selection as described by Goldberg and Deb (1991). The fittest individual in a pair is selected with probability 0.75. Otherwise the weaker individual is selected. |
Default: The original selection method described by Holland (1975), Selection_model = IMSLS_ROULETTE_WITH.
Output Keywords
N_generations—A scalar long indicating the number of generations used to find the fittest individual.
On_line_performance—An array of length N_generations containing on-line performance statistics for each generation.
Off_line_performance—An array of length N_generations containing off-line performance statistics for each generation.
Velocity—An array of length N_generations containing velocity statistics for each generation. The velocity for the ith generation is equal to 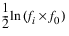 where fi is the maximum fitness for the ith generation.
where fi is the maximum fitness for the ith generation.
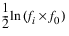 where fi is the maximum fitness for the ith generation.
where fi is the maximum fitness for the ith generation.Gen_statistics—A 2D array of size N_generations by 4 containing the maximum fitness, minimum fitness, average fitness and standard deviation of the fitness for each generation. The ith row of Gen_statistics contains the statistics for the ith generation. When N_generations<Max_generations, rows greater than N_generations – 1 are filled with NaN values. The four columns contain the following statistics calculated for each generation:
Last_generation—The last generation produced by the genetic algorithm.
Discussion
Genetic algorithms search for the optimum individual in a population. This is defined as the individual with the highest fitness. GENETIC_ALGORITHM returns the fittest individual in the last generation. Mathematically, this is equivalent to finding the values of the phenotypes that maximize a user-provided fitness function. Although there are no requirements that the fitness function be non-negative, in general, convergence to optimum fitness is faster when values of the fitness function are non-negative. Constraints can be applied by incorporating a penalty function within the fitness calculation. Phenotypes can consist of any combination of nominal, binary, integer and real values. Integer and real values must be encoded into a binary representation. This procedure provides for either Base-2 or Gray encoding. However, users can supply other encodings within the fitness function.
The function GENETIC_ALGORITHM uses the population data structure and fitness with simulated genetic processes of reproduction to search for the optimum individual, i.e. settings of phenotype values. Genetic algorithms have been successfully applied to a wide variety of optimization and search problems, see Holland (1975) and Goldberg (1985).
There are many refinements to the basic genetic algorithm originally described by Holland (1975). His basic algorithm begins with an initial population of n individuals, a fitness function, and probabilities for crossover and mutation of Crossover_prob and Mutate_prob respectively. The initial population is transformed from one generation to the next using the following steps:
1. Select n individuals from the current population to generate a mating pool.
2. Apply crossover with probability Crossover_prob to pairs of the selected individuals within the mating pool to produce two offspring.
3. Apply mutation with probability Mutate_prob to the offspring to generate the next generation.
4. Check stopping criteria. If they are met, stop and report the fittest individual within the last generation.
By default Holland’s approach to these steps are used. However, many variations of these can be selected using keywords.
The initial population can be generated automatically using the GA_RANDOM_POPULATION Function or it can be created by first creating individuals using the GA_INDIVIDUAL Function and then a population for those individuals using the GA_POPULATION Function.
By default Holland’s roulette wheel with replacement is used for selecting the mating pool. The Selection_model keyword allows users to select alternate selection methods including remainder, tournament and stochastic universal selection. Default crossover and mutation probabilities are Crossover_prob= 0.8 and Mutate_prob= 0.005. These defaults can be changed using the Crossover_prob and Mutate_prob keywords.
In the original algorithm only a single crossover point was randomly selected. The N_crossover_pts keyword allows users to designate any number of crossover points.
Standard crossover proceeds by combining the genes from both parents in the order found in those parents. Inversion crossover inverts this order for one of the parents. Inversion crossover is selected using the Invert_Crossover keyword.
For certain problems, such as the traveling salesman problem, standard crossover can produce infeasible individuals. One approach is to assign zero fitness to those solutions, but this can be very inefficient. Partially matched crossover is an approach that ensures individuals are feasible for a certain class of problems. If the problem is best represented using nominal phenotypes with values 0, 1, …, N_nominal – 1 where all values must appear once and only once in the chromosome, then partially matched crossover preserves that condition. Partially matched crossover is selected using the Pmx_crossover keyword.
One issue with some applications of genetic algorithms is premature convergence or convergence to false local solutions. This can occur when dominant individuals within early generations take over the population prematurely reducing population diversity. One approach to this problem is fitness scaling. This implementation allows users to use either linear or sigma fitness scaling. By default, no scaling is used. However, the Linear_scaling and Sigma_scaling keywords allow users to have fitness values automatically scaled before selection.
The genetic algorithm is stopped when any one of the stopping criteria is met. The algorithm is stopped when the number of generations exceeds Max_generations or when the maximum fitness exceeds Max_fitness. By default Max_generations = 100; this can be changed using Max_generations. By default Max_fitness is MACHINE(POS_INF); this can be changed using Max_fitness.
Example 1
De Jong (1975) examined the performance of a genetic algorithm for finding the maximum of a multivariate function. This is an example of optimizing a variation of De Jong’s R2 function:
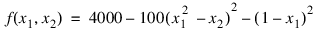
where:
–2.048 ≤ x1 ≤ 2.048 and –2.048 ≤ x2 ≤ 2.048
Since there were only two real phenotypes and the function is easily calculated, the phenotypes were encoded using discretization with 65,536 values over the interval [-2.048, 2.048]. By default, encoding and decoding is done within GENETIC_ALGORITHM. This allows the fitness function to calculate individual fitness using the real phenotypes instead of the chromosome. Both the chromosome and its phenotype representation are available within the individual data structure argument.
The default selection algorithm IMSLS_ROULETTE_WITH was used, but the number of crossover probability was set to 0.6. The genetic algorithm was more efficient using a lower crossover probability and Gray encoding instead of the defaults 0.7 and Base-2 encoding. Each generation consisted of 40 individuals.
PRO genetic_algorithm_ex1
@CMAST_COMMON
n = 40 ; Population size
n_generations = 0 ; Final number of generations
n_real = 2 ; Number of real phenotypes
r_intervals = [65536, 65536]
r_bounds = [-2.048, 2.048, -2.048, 2.048]
dashes= $
"**********************************************************"
RANDOMOPT, set=12345
chromosome = GA_CHROMOSOME(N_real=n_real, $
R_intervals=r_intervals, $
R_bounds=r_bounds)
population = GA_RANDOM_POPULATION(n, chromosome, $
/Gray_encoding, $
Fitness_fcn="deJongR2")
best_individual = GENETIC_ALGORITHM("deJongR2", population, $Print_level=IMSLS_FINAL, $
Max_fitness=3999.999, $
Crossover_prob=0.6, $
/Gray_encoding, $
Max_generations=1000, $
N_generations=n_generations, $
Gen_statistics=genStats, $
Last_generation=last_generation)
PRINT, $
"*****************GENERATION STATISTICS*****************"
PRINT,"Generation Max. Fit. Avg. Fit. Min. Fit. CV"
PRINT,"****************************************************"
gens = STRTRIM(SINDGEN((n_generations>last_generation.n)),2)
gens(0:9) = STRTRIM('0'+gens(0:9),2)FOR i=0L, n_generations-1 DO BEGIN
PRINT,"Gen. ",gens(i), " ", $
STRING(genStats(i,0),Format="(f11.5)")," ", $
STRING(genstats(i,2),Format="(f10.2)")," ", $
STRING(genstats(i,1),Format="(f10.2)")," ", $
STRING((100*genstats(i,3))/genStats(i,2),$
Format="(f9.2)")
ENDFOR
PRINT,""
PRINT,"LAST GENERATION"
PRINT,dashes
PRINT,"Indv Fitness ",$
"Chromosome X1 X2"
FOR i=0L, last_generation.n-1 DO BEGIN
PRINT,gens(i)," ", $
STRING(last_generation.fitness(i), $
Format="(f7.2)")," ",$
STRJOIN(STRTRIM $
(last_generation.individual(i).chromosome.allele, $
2),''),$
STRING(last_generation.individual(i).real_Phenotype(0),$
Format="(f7.3)")," ",$
STRING(last_generation.individual(i).real_Phenotype(1),$
Format="(f6.3)")
ENDFOR
PRINT,""
PRINT,"Maximum: ",$
STRING(last_generation.maxFitness,Format="(f7.2)"),$
" for individual ", STRTRIM(last_generation.indexFittest,2)
PRINT,"Minimum: ",$
STRING(last_generation.minFitness,Format="(f7.2)"),$
" for individual ", STRTRIM(last_generation.indexWeakest,2)
PRINT,"Average: ",$
STRING(last_generation.avgFitness,Format="(f7.2)")
PRINT,"Std. Dev: ",$
STRING(last_generation.stdFitness,Format="(f5.2)")
PRINT,dashes
END
;**************************************************************
; De Jong's R2 Function
;**************************************************************
FUNCTION deJongR2, individual
x1 = individual.real_Phenotype(0)
x2 = individual.real_Phenotype(1)
f = 100*(x1*x1-x2)*(x1*x1-x2) + (1.0-x1)*(1.0-x1)
f = 4000 - f
RETURN, f
END
Output
In this example, the print level is set to IMSLS_FINAL in order to print the optimum solution. The generation statistics are requested using the Gen_statistics keyword, and the last population is requested using the Last_generation keyword.
Although the maximum number of generations is set to 100 using the Max_generations option, the genetic algorithm halted after 26 generations when the maximum population fitness exceeded 3999.999.
OPTIMUM SOLUTION
Fitness: 3999.999512
Phenotypes:
Real: 2
Function Calculations: 1080
Population Size: 40
Number of Generations: 26
Real Phenotype(s):
1.023594 1.047844
Chromosome (Gray Encoded):
11100000000001011010000111000011
*****************GENERATION STATISTICS*****************
Generation Max. Fit. Avg. Fit. Min. Fit. CV
*******************************************************
Gen. 00 3996.12915 3244.16 578.92 28.62
Gen. 01 3996.25269 3725.90 2770.41 8.24
Gen. 02 3999.22974 3699.14 1917.26 10.28
Gen. 03 3999.22974 3683.41 2551.70 9.87
Gen. 04 3999.73779 3778.83 2551.70 8.35
Gen. 05 3999.73779 3823.50 3187.72 5.22
Gen. 06 3999.73779 3796.59 3187.72 5.76
Gen. 07 3999.73779 3850.94 3302.26 4.28
Gen. 08 3999.73779 3860.17 3358.92 3.93
Gen. 09 3999.73779 3886.33 3138.96 4.23
Gen. 10 3999.74683 3896.13 3292.64 3.85
Gen. 11 3999.74683 3900.24 3638.17 2.93
Gen. 12 3999.74683 3899.95 3376.35 3.17
Gen. 13 3999.74683 3900.57 3476.12 3.19
Gen. 14 3999.74683 3897.88 3408.28 3.36
Gen. 15 3999.74683 3908.28 2331.26 3.36
Gen. 16 3999.99585 3897.28 3301.18 3.94
Gen. 17 3999.99585 3910.99 3236.92 3.31
Gen. 18 3999.99585 3953.46 3429.17 1.31
Gen. 19 3999.99585 3944.98 3764.08 1.41
Gen. 20 3999.99585 3945.62 3751.01 1.41
Gen. 21 3999.99585 3934.07 3751.10 1.81
Gen. 22 3999.99585 3947.08 3739.52 1.62
Gen. 23 3999.99609 3943.99 3652.86 1.95
Gen. 24 3999.99609 3942.42 3652.86 1.99
Gen. 25 3999.99927 3970.69 3845.42 0.92
LAST GENERATION
**************************************************************
Indv Fitness Chromosome X1 X2
00 3916.58 11100000000011001100110001010011 1.023 0.134
01 3996.09 01010000000001011100110001010011 -0.512 0.134
02 3965.22 11010000000011101110111110011011 0.511 0.849
03 3993.47 01100101000001011010000001100011 -0.928 1.028
04 3941.02 11010000000001011010000001100011 0.512 1.028
05 3998.11 11110000000001011100110001110011 0.512 0.134
06 3951.04 01100100000101011100110001100011 -0.898 0.132
07 3919.76 01000000000001011110110001110011 -0.000 0.890
08 3965.35 11110000000001011110111110011011 0.512 0.849
09 3913.56 11010000000011101010111101110011 0.511 1.190
10 3948.39 11010000000011101110001110110011 0.511 0.978
11 3996.04 01010000000001011100110001100011 -0.512 0.132
12 3997.44 11100000100111001110111101101001 1.009 0.859
13 3992.45 11110000000001110100000001000011 0.512 -0.008
14 3958.07 01010000000101011110110001110011 -0.510 0.890
15 3951.33 01100100000001011100110001100011 -0.896 0.132
16 3948.39 11010000000011101110001110110011 0.511 0.978
17 4000.00 11100000000001011010000111010001 1.024 1.046
18 3999.96 11100000000001011010000001100011 1.024 1.028
19 3925.31 01010110010001011010000111010001 -0.440 1.046
20 3999.37 11010000000001011101111001101111 0.512 0.325
21 3845.42 01100101010110110101110001100011 -0.921 -0.380
22 3999.64 11010101000011101100111001000011 0.415 0.184
23 3992.37 01100000000011101110111101101001 -1.023 0.859
24 3990.55 01010000000001110100000001110011 -0.512 -0.006
25 3993.26 11110000000011101100000001000011 0.513 0.008
26 3964.15 11010000000011101110111101110011 0.511 0.858
27 3998.36 11010000000011101100110010011011 0.511 0.143
28 3950.63 01100101000011101100111001000011 -0.927 0.184
29 3996.08 01010000000001011100110001110011 -0.512 0.134
30 3997.89 01010110000011101100111001100111 -0.447 0.188
31 3998.11 11110000000001011100110001110011 0.512 0.134
32 3993.47 01100101000001011010000001100011 -0.928 1.028
33 3992.45 11110000000001110100000001000011 0.512 -0.008
34 3916.61 11100000000011011100110001010011 1.023 0.134
35 3929.38 01010110000001011010000001100011 -0.448 1.028
36 3997.89 01010110000011101100111001100011 -0.447 0.188
37 4000.00 11100000000001011010000111010001 1.024 1.046
38 4000.00 11100000000001011010000111000011 1.024 1.048
39 3991.28 01100100000001011010000001110001 -0.896 1.030
Maximum: 4000.00 for individual 38
Minimum: 3845.42 for individual 21
Average: 3970.61
Std. Dev: 35.71
**************************************************************
Example 2
The traveling saleman problem creates a problem for traditional crossover. In this problem, the objective is to find the shortest route while traveling to each city once. In this example, there are eight cities, labeled using the letters a-h, with distances ranging from 17 to 113 miles.
Traditional crossover would create unfeasible routes; that is some routes after crossover would not visit every city once. Some would not be visited and others would be visited more than once.
Partially matched crossover (PMX) preserves the feasibility of a route. In the general sense, PMX assumes that the nominal phenotypes consists of a string of numbers from zero to N_nominal – 1, with each number appearing once and only once in that string. Partially matched crossover uses two crossover points within the nominal portion of the chromosome and swaps the middle segment between the parents. The first and third segments are manipulated to ensure the resulting offspring is feasible.
PRO t_gen_alg_ex2
@CMAST_COMMON
DECLARE FUNC, pmxFitness
n = 50 ; population size
n_nominal = 8 ; number of nominal phenotypes
n_categories = [8, 8, 8, 8, $
8, 8, 8, 8] ; nominal category limits
cities = ["a", "b", "c", "d", $ ; Cities Label a-h
"e", "f", "g", "h"]
RANDOMOPT,set=54321
chromosome = GA_CHROMOSOME(N_nominal=n_nominal, $
N_categories=n_categories)
population = GA_RANDOM_POPULATION(n, chromosome,$
/Pmx_crossover,$
Fitness_fcn="pmxFitness")
best_individual = GENETIC_ALGORITHM( $
"pmxFitness", population, $
Print_level=IMSLS_FINAL, $
/Pmx_crossover, $
/Invert_crossover, $
Crossover_prob=0.8, $
Max_generations=10, $
Gen_statistics=genStats, $
N_generations=n_generations, $
Last_generation=last_generation)
PRINT,"GENERATION STATISTICS"
PRINT,"Total Number of Generations: ",$
STRTRIM(n_generations,2)
PRINT,""
PRINT, $
"*****************GENERATION STATISTICS*****************"
PRINT,"Generation Max. Fit. Avg. Fit. Min. Fit. CV"
PRINT,"****************************************************"
gens = STRTRIM(SINDGEN(last_generation.n),2)
gens(0:9) = STRTRIM('0'+gens(0:9),2)FOR i=0L, n_generations-1 DO BEGIN
PRINT,"Gen. ",gens(i), " ", $
STRING(genStats(i,0),Format="(f10.2)")," ", $
STRING(genstats(i,2),Format="(f10.2)")," ", $
STRING(genstats(i,1),Format="(f10.2)")," ", $
STRING((100*genstats(i,3))/genStats(i,2),$
Format="(f9.2)")
ENDFOR
PRINT,""
PRINT," LAST GENERATION"
PRINT,"*************************************"
PRINT,"Individual Fitness Phenotype Values "
avg = last_generation.avgFitness
FOR i=0L,last_generation.n-1 DO BEGIN
x1 = last_generation.fitness(i)
IF x1 EQ last_generation.maxFitness THEN $
stars = '****' $
ELSE stars = " "
PRINT,stars,gens(i)," ",$
STRTRIM(STRING(x1,Format="(I3)"),2)," ",$
cities(last_generation.individual(i).nominal_Phenotype)
ENDFOR
PRINT,""
PRINT,"Average Fitness: ", $
STRING(avg,Format="(f6.1)")
PRINT,"*************************************"
PRINT,"OPTIMUM SOLUTION:"
PRINT,""
PRINT,"Fitness: ", $
pmxFitness(best_individual)
PRINT,"Chromosome: ",$
STRTRIM(best_individual.chromosome.allele,2)
PRINT,"Phenotype Values: ",$
STRJOIN(cities(best_individual.nominal_Phenotype)," -> ")
END
;**************************************************************
; FITNESS FUNCTION
;**************************************************************
FUNCTION pmxFitness,individual
n_nominal = 8 ; number of nominal phenotypes
; cities: a b c d e f g h
distances =[[ 0, 17, 27, 73, 61, 57, 51, 23],$ ; a
[17, 0, 37, 73, 72, 74, 66, 40],$ ; b
[27, 37, 0, 48, 35, 49, 65, 50],$ ; c
[73, 73, 48, 0, 47, 82, 113, 95],$ ; d
[61, 72, 35, 47, 0, 38, 80, 78],$ ; e
[57, 74, 49, 82, 38, 0, 48, 65],$ ; f
[51, 66, 65, 113, 80, 48, 0, 40],$ ; g
[23, 40, 50, 95, 78, 65, 40, 0]] ; h
n_nominal = individual.chromosome.n_nominal
k = individual.chromosome.nominalIndex+1
f = 0.0
FOR i=k,(k+n_nominal-1)-1 DO BEGIN
i1 = individual.nominal_Phenotype(i-1)
i2 = individual.nominal_Phenotype(i)
f = f + distances(i1,i2)
ENDFOR
RETURN, 516.-f
END
Output
This program produced the following output. Since the print level was set to IMSLS_FINAL, the optimum solution was printed. The generation statistics were requested using the Gen_statistics keyword, and the last population was requested using the Last_generation keyword.
The maximum number of generations was set to 10. The genetic algorithm found the optimum route after evaluating the fitness of 550 routes in 10 generations. Generation zero is the initial generation provided to the algorithm and is not counted towards the maximum generation count.
OPTIMUM SOLUTION
Fitness: 259.000000
Phenotypes:
Nominal: 8
Function Calculations: 550
Population Size: 50
Number of Generations: 10
Nominal Phenotype(s):
6 7 1 0 2 3 4 5
Chromosome (Base-2 Encoded):
6 7 1 0 2 3 4 5
GENERATION STATISTICS
Total Number of Generations: 10
*****************GENERATION STATISTICS*****************
Generation Max. Fit. Avg. Fit. Min. Fit. CV
*******************************************************
Gen. 00 209.00 110.22 9.00 43.49
Gen. 01 209.00 136.38 10.00 27.83
Gen. 02 230.00 142.00 56.00 26.60
Gen. 03 233.00 150.76 34.00 33.75
Gen. 04 258.00 147.76 33.00 34.75
Gen. 05 259.00 159.98 29.00 34.16
Gen. 06 259.00 158.80 60.00 31.44
Gen. 07 259.00 145.84 42.00 27.25
Gen. 08 259.00 139.86 66.00 28.04
Gen. 09 259.00 143.38 28.00 28.45
LAST GENERATION
*************************************
Individual Fitness Phenotype Values
00 180 a b g h e f c d
01 143 h c b g f e a d
02 78 f a g b e d h c
03 131 g a f d c e b h
04 93 a f e g c d h b
05 174 a b h g e c d f
06 93 f h d e b g a c
07 128 f a h g d c e b
08 104 g d h c a b e f
09 127 e f a c d g b h
10 138 g f a h d c e b
11 152 f d e g c h a b
12 211 f a h g b c e d
13 115 h a b c f d g e
14 158 g b a c h e f d
15 105 c b g f a e d h
16 178 g f b c d e a h
17 139 f h g b c e a d
18 186 d c a h b f e g
19 105 c b g f a e d h
****20 259 g h b a c d e f
21 107 g c d b f h a e
22 107 e a h f b d c g
23 136 d h a c f g b e
24 139 c a g h f b d e
25 181 a f e d c g h b
26 161 d e h b g f c a
27 107 g f b d h a e c
28 115 h a b c f d g e
29 190 e f d c h b a g
30 162 e f d b c h a g
31 119 d a f g b h e c
32 135 b h f e d g a c
33 135 b h f e d g a c
34 224 e f d c a b h g
35 111 e g d c h b a f
36 152 g a c d f e h b
****37 259 g h b a c d e f
38 106 a e b g f h c d
39 146 d a b e f h g c
40 58 c f a h e b g d
41 188 f a b c h g e d
42 91 e h b c f a g d
43 132 f b d c g h a e
44 136 f a b g c h e d
45 109 g e f a c b d h
46 173 g f h e c a b d
47 93 c e h g b f a d
48 142 g e d c b h f a
49 91 g c d h f b a e
Average Fitness: 140.0
*************************************
OPTIMUM SOLUTION:
Fitness: 259.000
Chromosome: 6 7 1 0 2 3 4 5
Phenotype Values: g -> h -> b -> a -> c -> d -> e -> f
Example 3
This example uses the N-Queens problem to illustrate the use of a fitness function with parameters in implementing a genetic algorithm. The N-Queens problem is derived from chess. The genetic algorithm provides an efficient search for a valid solution. For this problem the chess board consists of N rows and N columns. The objective is to place N queens on this board with no conflicts. A conflict occurs when one queen can move and capture another. Since queens can move diagonally, vertically and horizontally this problem is challenging when N becomes large.
One solution for N = 4 is displayed in Table 14-7: N-Queens Solution. A valid solution must place every queen in a different row and column. The problem is to ensure the queens are not in conflict because of lying on the same diagonal.
Row/Col | 0 | 1 | 2 | 3 |
|---|---|---|---|---|
0 | Q | |||
1 | Q | |||
2 | Q | |||
3 | Q |
Similar to the traveling salesman problem, the N-Queens problem can be expressed using N nominal phenotypes with values 0, 1, …, N–1. The value of the ith phenotype represents the row number for the queen in the ith column. This ensures that any arrangement of the phenotype values represents a board with N queens, each placed in a different row and column.
The solution for N queens displayed above can be represented by the phenotype values {1, 3, 0, 2}. The search looks for arrangements that also do not place queens on the same diagonal. Two queens fall on the same diagonal if the absolute value of the difference between their row numbers equals the absolute value of the difference between their column numbers.
This example uses this representation with a fitness function equal to (N – C) where C equals the number of conflict among the queens. With this fitness, a solution to the N-Queens has a fitness of N.
PRO gen_alg_ex3
DECLARE FUNC, queensFitness
@CMAST_COMMON ; define IMSLS constants
; COMMON block for data we will pass to the fitness
; function.
COMMON EX3fitness_args, n_queens
n = 500 ; population size
n_queens = 12 ; number of nominal phenotypes
n_categories = LONARR(n_queens)
RANDOMOPT,set=54321
maxFit = n_queens - 0.5
n_categories(*) = n_queens
chromosome = GA_CHROMOSOME( $
N_nominal=n_queens,$
N_categories=n_categories)
population = GA_RANDOM_POPULATION(n, $
chromosome, $
/Pmx_crossover, $
Fitness_fcn="queensFitness")
best_individual = GENETIC_ALGORITHM( $
"queensFitness", population, $
Print_level=IMSLS_FINAL, $
/Pmx_crossover, $
Linear_scaling=2.0, $
Crossover_prob=0.7, $
Mutate_prob=0.01, $
Max_generations=10000, $
Max_fitness=maxFit, $
Gen_statistics=genStats, $
N_generations=n_generations)
PRINT,"GENERATION STATISTICS"
PRINT,"Total Number of Generations: ",$
STRTRIM(n_generations,2)
PRINT,"Generation Max. Fit. Min. Fit. Avg. Fit. CV"
PRINT,"*************************************************"
gens = STRTRIM(SINDGEN(n_generations),2)
IF n_generations GT 9 THEN $
gens(0:9) = STRTRIM('0'+gens(0:9),2)FOR i=0L, n_generations-1 DO BEGIN
PRINT,"Gen. ",gens(i), " ", $
STRING(genStats(i,0),Format="(f11.5)")," ", $
STRING(genstats(i,1),Format="(f10.2)")," ", $
STRING(genstats(i,2),Format="(f10.2)")," ", $
STRING((100*genstats(i,3))/genStats(i,2),$
Format="(f9.2)")
ENDFOR
PRINT,"*************************************************"
PRINT,""
PRINT,"OPTIMUM SOLUTION:"
PRINT,""
PRINT,"Fitness: ", $
queensFitness(best_individual)
PRINT,"Chromosome: ",$
STRTRIM(best_individual.chromosome.allele,2)
dashes = STRJOIN([REPLICATE("--",n_queens-1),'---'],'')PRINT,"Board Positions: "
FOR i=0L, n_queens-1 DO BEGIN
line = REPLICATE("| ",n_queens+1)ret = WHERE( $
best_individual.nominal_Phenotype EQ i)
IF ret(0) NE -1 THEN line(ret) = "|Q"
PRINT, dashes
PRINT, STRJOIN(line,"")
ENDFOR
PRINT, dashes
END
FUNCTION queensFitness, individual
COMMON EX3fitness_args, n_queens
f = 0.0 ; Fitness value
FOR i=0L, n_queens-2 DO BEGIN
FOR j=i+1, n_queens-1 DO BEGIN
k = individual.chromosome.allele(i) - $
individual.chromosome.allele(j)
IF ABS(i-j) EQ ABS(k) THEN f = f + 1.0
ENDFOR
ENDFOR
f = n_queens - f
RETURN, f
END
Output
This program produced the following solution to the N-Queens problem with N=50 queens. Notice that some of the minimum fitness values are negative. This alters the random selection of the fittest parents, but if these values are few and small, then the effect is not enough to halt the genetic algorithm.
For 50 queens there are over 1064 ways of placing queens ensuring each is in its own row and column. An exhaustive search of these possible solutions to find a arrangement without diagonal conflicts would be time consuming. The genetic algorithm search found a solution in only 568 generations, requiring 284,500 function evaluations.
OPTIMUM SOLUTION
Fitness: 12.000000
Phenotypes:
Nominal: 12
Function Calculations: 2500
Population Size: 500
Number of Generations: 4
Nominal Phenotype(s):
8 10 2 7 11 1 3 0 6 4 9 5
Chromosome (Base-2 Encoded):
8 10 2 7 11 1 3 0 6 4 9 5
GENERATION STATISTICS
Total Number of Generations: 4
Generation Max. Fit. Min. Fit. Avg. Fit. CV
*************************************************
Gen. 0 10.00000 -14.00 4.41 61.95
Gen. 1 10.00000 -3.00 4.77 51.50
Gen. 2 10.00000 -5.00 5.10 46.84
Gen. 3 12.00000 -4.00 5.31 42.44
*************************************************
OPTIMUM SOLUTION:
Fitness: 12.0000
Chromosome: 8 10 2 7 11 1 3 0 6 4 9 5
Board Positions:
-------------------------
| | | | | | | |Q| | | | |
-------------------------
| | | | | |Q| | | | | | |
-------------------------
| | |Q| | | | | | | | | |
-------------------------
| | | | | | |Q| | | | | |
-------------------------
| | | | | | | | | |Q| | |
-------------------------
| | | | | | | | | | | |Q|
-------------------------
| | | | | | | | |Q| | | |
-------------------------
| | | |Q| | | | | | | | |
-------------------------
|Q| | | | | | | | | | | |
-------------------------
| | | | | | | | | | |Q| |
-------------------------
| |Q| | | | | | | | | | |
-------------------------
| | | | |Q| | | | | | | |
-------------------------